Our Services
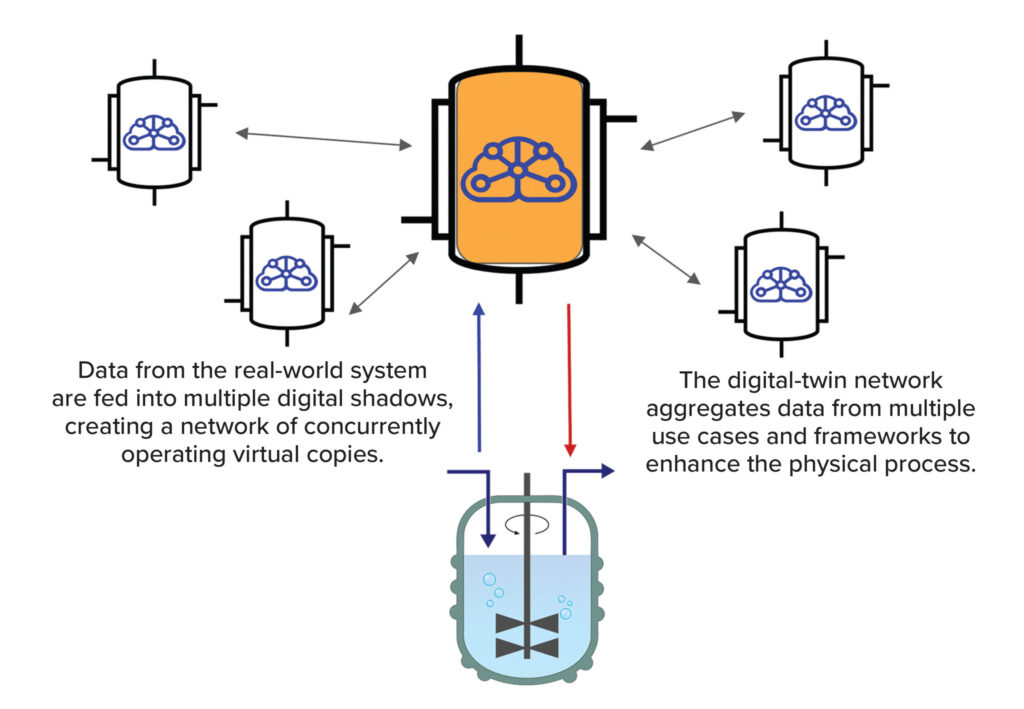
Divergene collaborates closely with CROs, CDMOs, cell & gene therapy (CGT) companies to enhance their next-generation sequencing capabilities. Our support spans from upstream vector engineering to downstream vector QC. We also utilize AI driven digital-twin network aggregates data from multiple use cases and frameworks to enhance the physical process. Our AI driven digital-twin emulates the complexity of biological processes and addresses all of the difficulties related to the continuous variability associated with biopharmaceutical operations.

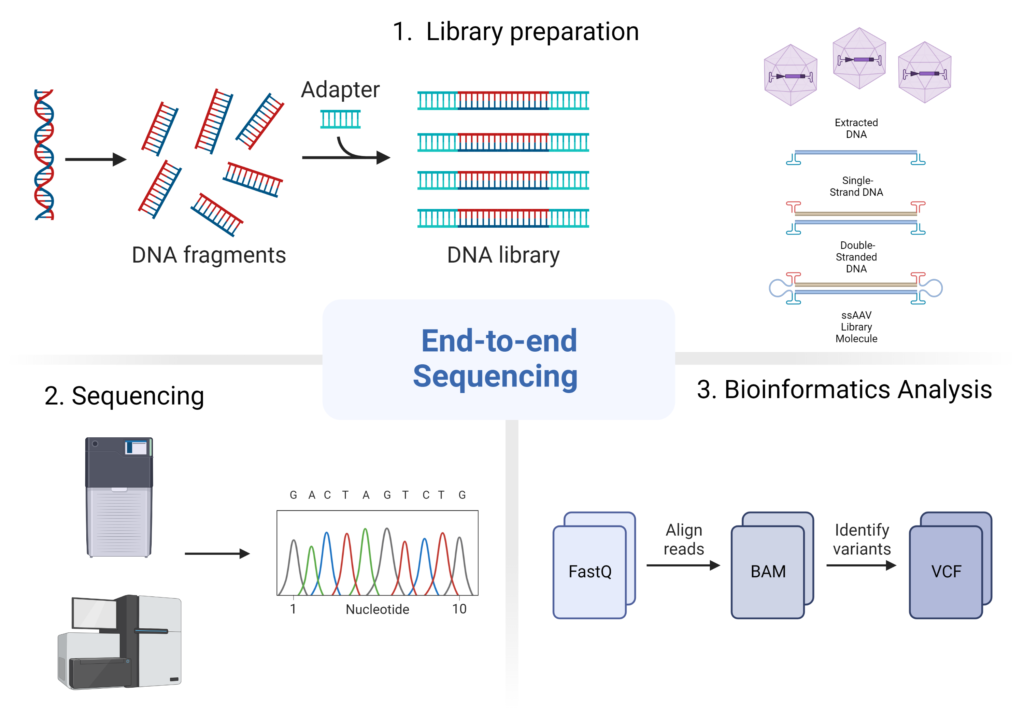
End-to-end Sequencing
Whether you are looking for AAV capsid library enrichment or interested in post-infection solutions, Divergene utilizes the latest technologies on the market, including PacBio® long-read (>10kb) and Illumina® short-read sequencing, to provide custom, in-depth solutions for complex AAV gene therapy applications.
- AAV Genome Sequencing
- Vector Discovery
- Confirm Plasmid Sequences
- Quantify AAV Expression
- Validate Packaged Material
- Capsid Engineering
- Transcriptome Analysis
If any of the applications interest you, get in touch with us.
Bioreactor and Bioprocess Simulation by AI in 3 Approaches
No Connection in Real Time: The models present a static picture of how a physical system operated at a specific time. We can understand factors and values that determine the model’s broader theoretical framework.
One-Way Connection: A one-way in silico replica can be useful for applying process analytical technology (PAT), real-time monitoring, and remotely initiated process control — all of which can improve an ongoing process.
Full Connection: Bidirectional connectivity enables a model to support a physical manufacturing system in a growing number of ways. Real-time updates to the model provide for such activities as intelligent closed-loop control. Thus, batches can be produced in the right way during every iteration through continuous monitoring and “smart” operation.
Why Choose Us
Superior Speed & Data Quality
Our robust framework, backed by stringent quality control measures, consistently delivers high-quality data.
Ph.D.-Level Support
Our experienced team is dedicated to offering rapid insights and responses throughout the entire process, ensuring quick turnaround times.
Customized solutions
We provide comprehensive support across all lab stages, ensuring a smooth transition from wet lab to dry lab. Our services include actionable dashboards and reports to enhance results and insights.
cell & gene therapy Expert
We are the trusted partner of choice for leading cell and gene therapy biotechs and biopharma companies.